1. 引言
基因调控网络是由一系列相互作用的生物分子组成的动态演化系统 [1] 。每一个网络节点代表一种生物分子,而网络边则表示分子间的相互作用(包括抑制作用和激活作用)。调控网络各节点的表达状态受相互作用的影响而不断变化,这种顺序变化的状态序列是控制细胞内各种生命活动的控制信号。在实际演化环境中,调控网络能够克服生物化学过程中的各种随机干扰,稳定地保持状态演化的稳定性。因此,研究调控网络的状态演化规律及其影响因素是系统生物学的重要议题 [2] - [5] 。
前期研究表明,调控网络的拓扑结构与网络的稳定性特征有密切联系。Li等人 [2] 运用经典的布尔模型对调控网络的状态演化过程进行建模,首次发现了调控网络的状态稳定性和路径稳定性,同时,该结果也指示了网络拓扑结构是产生这些稳定性特征的决定性因素。Wang等人 [5] 和Zhang等人 [6] 进一步研究了如何利用网络的稳定性特征来提高逆向学习网络结构的效率。Wu等人 [7] 提出了调控熵来描述调控网络的结构特征。调控熵是指网络内部节点之间的调控关系的一致性。通过模拟数据和实际网络验证发现,调控网络的稳定性特征与其调控熵大小有显著的相关性。
为了准确定位产生网络稳定性的特征结构模块,Zhang等人 [8] 从链式网络模型出发,建立了调控网络的主干结构模型—理想传递链(Ideal Transmission Chain, ITC)模型,并首次从理论上证明了ITC模型是产生状态稳定性的充分和必要条件。此外,真实调控网络的实例研究也表明,ITC模型对其状态稳定性具有决定性作用。
本文在文献 [8] 的基础进一步讨论影响网络路径稳定性的结构因素。真实的调控网络是以ITC模型为基本主干,同时包含多个分支链路的树状结构,而且,分支链路的结构基本相同,只是接入ITC模型的位置不同。因此,本文研究了三种典型接入位置,即在ITC模型之外,在ITC模型的起点以及在ITC模型的末端接入分支链路。针对这三种情况,分别证明了加入分支链路后路径稳定性的变化情况。最后,用实验模拟的方式研究了在其他位置加入分支链路后路径稳定性的变化情况。
2. 方法
2.1. 布尔网络模型简介
本研究使用布尔网络模型来描述基因调控网络的状态演化过程。布尔网络模型(Boolean network model)是一种常用的网络建模方法 [2] [5] [9] ,它用0~1取值的布尔变量来表示网络节点的状态。网络节点的状态取0,对应实际情况下该种基因不表达或低表达;节点取值为1表示该基因高表达。节点之间的作用边也被简化成抑制和激活两种类型,分别取权值+1和−λ两种情况。λ体现了抑制作用对激活作用的相对权重:λ = 1表示抑制作用和激活作用等权;另一种情况是λ = ∞,表示抑制作用具有决定性作用。后一种模型也成为一票否决模型。在本文的研究中,我们主要考虑一票否决模型。
在布尔模型下,各节点的状态演化是同步进行的,具体规则如下:对于包含n个节点的网络,假定在t时刻节点i的状态是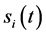 ,那么,在t + 1时刻,有
,那么,在t + 1时刻,有

其中, 是节点j到节点i的作用边权值:如果是激活边,
是节点j到节点i的作用边权值:如果是激活边, ;是抑制边的话,
;是抑制边的话,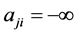 。我们将同一时刻(如t时刻)网络各个节点的状态组合成n维的二值向量,即
。我们将同一时刻(如t时刻)网络各个节点的状态组合成n维的二值向量,即 ,该向量就表示网络在该时刻的状态。
,该向量就表示网络在该时刻的状态。
在布尔网络模型的框架下,调控网络内部的各种生物分子之间的调控关系被简化为相应网络节点之间的连接关系(用连接关系矩阵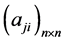 表示)。这种连接关系可以用网络拓扑图表示(如下面的图1所示)。布尔网络简单而有效,能够较完整地刻画调控网络的整体动力学行为特征 [2] ,因此,在布尔模型的基础上,调控网络的研究重点就是网络的拓扑关系对调控网络动力学性质的影响。
表示)。这种连接关系可以用网络拓扑图表示(如下面的图1所示)。布尔网络简单而有效,能够较完整地刻画调控网络的整体动力学行为特征 [2] ,因此,在布尔模型的基础上,调控网络的研究重点就是网络的拓扑关系对调控网络动力学性质的影响。
2.2. 路径稳定性简介
路径稳定性表示网络状态演化路径的重叠性。文献 [2] 引入了W值来表示路径稳定性。具体计算思路如下:假设网络从状态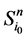 开始,经Li步到达演化终点
开始,经Li步到达演化终点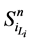 ,每一步路径上的路径通量(定义为通过该路径的所有网络状态的数目)记作
,每一步路径上的路径通量(定义为通过该路径的所有网络状态的数目)记作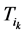 ,
,
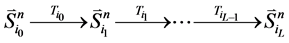
那么,对于状态点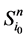 的
的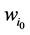 值为
值为
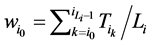
进一步,网络整体的W值是各个状态w值的平均,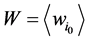 。W值体现了网络中各个状态路径的平均重合度。但是,这种对路径稳定性的定义方法容易受到网络规模的影响,因为节点数目多的网络拥有更多的网络状态数,因此,其状态空间中每一条演化路径上的路径通量会偏大。从而导致节点数目多的网络W值越大。
。W值体现了网络中各个状态路径的平均重合度。但是,这种对路径稳定性的定义方法容易受到网络规模的影响,因为节点数目多的网络拥有更多的网络状态数,因此,其状态空间中每一条演化路径上的路径通量会偏大。从而导致节点数目多的网络W值越大。
为了消除由于网络规模不同导致的网络状态数目的变化及其对路径稳定性的影响,本研究中采用相对重合度来表示路径稳定性,即定义W值为
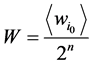
 (a)
(a) 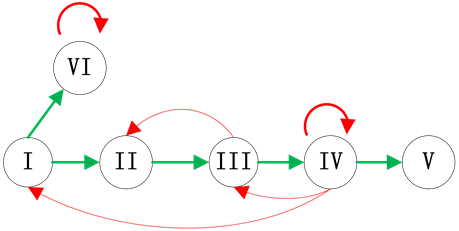 (b)
(b)  (c)
(c)
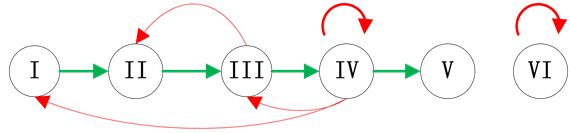
Figure 1. Examples of branches in ITC model; (a) Independent branch; (b) Branch at the start node; (c) Branch at the end node
图1. 分支链路位置示意图;(a) 独立分支链路;(b) 起点分支链路;(c) 末端分支链路
3. 结果
前期研究表明 [8] ,真实的调控网络是以ITC模型为基本主干,同时包含多个分支链路的树状结构。在本节中,我们将通过严格的论证来分析不同位置的分支结构对网络的路径稳定性的影响情况,揭示在真实调控网络中路径稳定性的形成机制。
结合典型的生物学特征,我们主要考虑了三种情况的分支链路,即独立链路(分支链路与ITC模块没有激活关系连接),起点分支链路(分支链路被调控网络的起始节点激活)和末端分支链路(分支链路被调控网络的最后一个传递节点激活)。在真实的调控网络中,这三种情况都有典型的例子:例如,Fission yeast调控网络中的SK节点就是独立链路;Budding yeast网络中的SBF节点和Cln1,2节点是起点分支链路;而Budding yeast网络中的Swi5节点就是末端分支链路。这三种分支链路的具体示意图如图1所示。对于在网络中其他位置插入分支链路,网络路径稳定性的变化情况将在模拟实验中给出。
3.1. 独立链路对ITC路径稳定性的影响
独立链路(如图1(a)所示)与ITC模块之间没有激活作用相联系,因此,该节点会在自身抑制作用的控制下被关闭。图2中对比了加入独立链路前后的网络状态演化路径的变化情况。
对于原网络(即加入链路之前)任意一条起点为 ,长度为
,长度为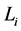 的演化路径,在加入独立链路之后,该路径上的每一个状态点都会派生出两个状态点,即
的演化路径,在加入独立链路之后,该路径上的每一个状态点都会派生出两个状态点,即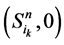 和
和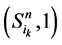 。它们分别对应着独立链路上的节点取0或者1。对于状态点
。它们分别对应着独立链路上的节点取0或者1。对于状态点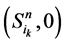 ,它们会保持原有的演化路径。而
,它们会保持原有的演化路径。而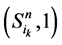 状态点会在推迟一个单位时间
状态点会在推迟一个单位时间
 (a)
(a)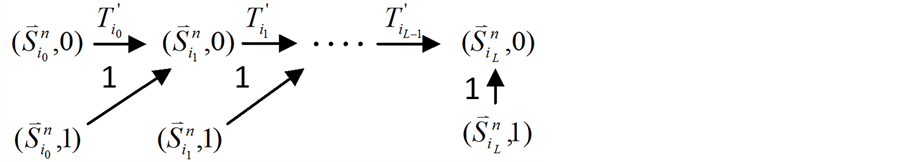 (b)
(b)
Figure 2. Analysis of the trajectory stability after inserting an independent branch; (a) Before inserting the branch, the trajectory of state evolution and the flow of each link; (b) After inserting the branch, the trajectory of state evolution and the flow of each link
图2. 独立链路对ITC模型的路径稳定性影响分析;(a) 加入独立链路之前,状态演化路径及各路径通量情况;(b) 加入独立链路之后,状态演化路径及各路径通量情况
汇入原有的演化路径(如图2所示)。对于这种情况,有如下的结论:
结论1:加入独立链路之后,ITC网络的W值降低。
证明:记 表示原有演化路径上第k步的路径通量,那么,加入独立链路之后该通量变为
表示原有演化路径上第k步的路径通量,那么,加入独立链路之后该通量变为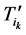

进一步,原有状态点 以及新的状态点
以及新的状态点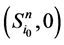 和
和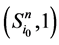 所对应的w值分别为
所对应的w值分别为
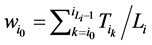
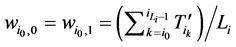 (1)
(1)
将公式(1)带入,即可以推导出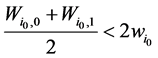 。
。
最后,加入独立链路点之后,网络的路径稳定性指标变成
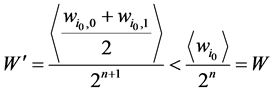 (2)
(2)
这表明加入独立链路之后,网络的W值下降了。
3.2. 起点分支链路对ITC路径稳定性的影响
接下来,考虑起点处的分支链路(如图1(b)所示),分析方法与独立链路的情况相同。
在加入分支链路之前,依据起始节点的0~1状态,所有的网络状态可以分成两类,即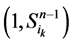 和
和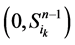 。这两类点的一般演化途径及各路径通量由图3(a)所示。加入分支链路之后,原有的两类状态派生出四类状态点(依据分支链路节点的0~1状态),即
。这两类点的一般演化途径及各路径通量由图3(a)所示。加入分支链路之后,原有的两类状态派生出四类状态点(依据分支链路节点的0~1状态),即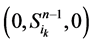 、
、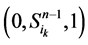 、
、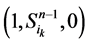 和
和 。这四类状态的演化路径及其通量由图3(b)所示。
。这四类状态的演化路径及其通量由图3(b)所示。
从图中可以看出,加入分支链路之后,两条从状态点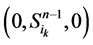 和状态点
和状态点 起始的演化路径替代了原来的主干路径,而其他的状态点(如
起始的演化路径替代了原来的主干路径,而其他的状态点(如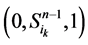 和
和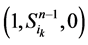 )都会延迟一个单位时间汇入干路径。而且,这两条主体路径在第k+1步(比原来推迟了一个单位时间)发生汇聚。由此分析,可以得出以下结论:
)都会延迟一个单位时间汇入干路径。而且,这两条主体路径在第k+1步(比原来推迟了一个单位时间)发生汇聚。由此分析,可以得出以下结论:
结论2:加入起点分支链路之后,网络的路径稳定性指标值会降低。
证明:具体的证明过程类似于独立链路的情形,在此省略。
3.3. 末端分支链路对ITC路径稳定性的影响
本小节考虑最后一种情况,即分支链路联接在ITC网络的最后一个传递节点上(如图1(c)所示)。
在没有加入分支链路之前,对于任意一条收敛路径(如图4(a)所示),其终点状态就是网络的全局吸引子。也就是说,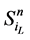 中仅有最后一个节点表达(取值为1),而其他所有的网络节点都被关闭(取值为0)。
中仅有最后一个节点表达(取值为1),而其他所有的网络节点都被关闭(取值为0)。
在加入分支链路之后,原路径上的每一个状态点都会对应成两个状态,分别为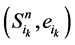 和
和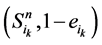 。其中,
。其中,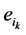 (取值0或者1)表示主路径上每一个时间点链路节点的状态。它们的演化路径如图4(b)所示。之所以这么标记是因为加入末端分支链路之后,原有的演化路径发生了变化,无法确定是由主路径上状态点中分支节点的具体取值。但是,可以肯定,无论
(取值0或者1)表示主路径上每一个时间点链路节点的状态。它们的演化路径如图4(b)所示。之所以这么标记是因为加入末端分支链路之后,原有的演化路径发生了变化,无法确定是由主路径上状态点中分支节点的具体取值。但是,可以肯定,无论 取值如何,其对应的状态
取值如何,其对应的状态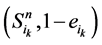 会延迟1个单位时间汇入主路径。
会延迟1个单位时间汇入主路径。
另一方面,在状态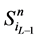 中,网络的最后一个传递节点(也就是分支链路接入的节点)必然是打开状态,这使得
中,网络的最后一个传递节点(也就是分支链路接入的节点)必然是打开状态,这使得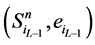 不会直接变成
不会直接变成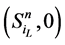 ,而是先经过
,而是先经过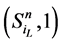 再到
再到 。这一现象说明,原来长度为
。这一现象说明,原来长度为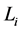 步的演化路径被增加了一个步长(
步的演化路径被增加了一个步长(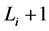 步)。这能够增加单个状态的W,从而提高网络整体的W值。
步)。这能够增加单个状态的W,从而提高网络整体的W值。
结论3:加入末端分支链路之后,网络的路径稳定性指标值会增加。
证明:加入末端分支链路之后,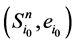 和
和 分别对应的W值为
分别对应的W值为
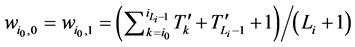 (3)
(3)
所以,对该状态点而言,加入分支链路后W值的增加量为
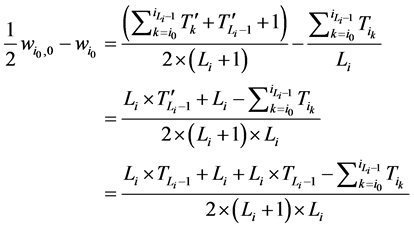
由于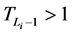 ,而且
,而且 ,故
,故
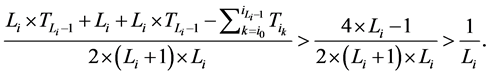
从而网络整体的W值也相应增加了。
3.4. 模拟实验结果
为了验证上述理论分析的结果,我们进行了随机模拟实验。对给定网络规模(即网络节点数目)的ITC模块,在不同的位置加入分支链路,并计算加入链路后的网络路径稳定性(W值)。典型的调控网络都包含10个左右的网络节点,我们的模拟实验将网络主干的节点数定在6~12个。给定节点数目,随机选择一个ITC链路构成网络主干,然后在指定位置上加入分支链路并计算W值。重复上述过程10,000次,计
 (a)
(a)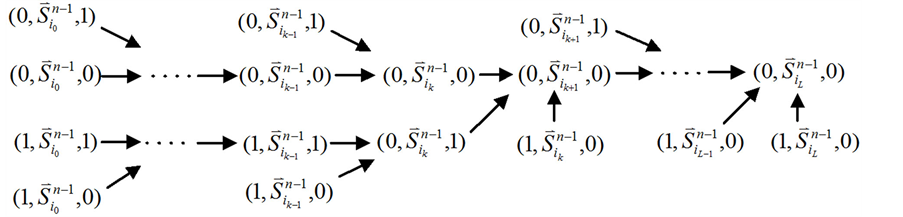 (b)
(b)
Figure 3. Analysis of the trajectory stability after inserting a branch at the start node; (a) Before inserting the branch, the trajectory of state evolution and the flow of each link; (b) After inserting the branch, the trajectory of state evolution and the flow of each link
图3. 起点分支链路对ITC模型的路径稳定性影响分析;(a) 加入起点分支链路之前,状态演化路径及各路径通量情况;(b) 加入起点分支链路之后,状态演化路径及各路径通量情况
 (a)
(a)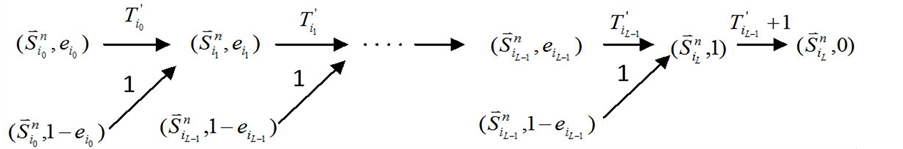 (b)
(b)
Figure 4. Analysis of the trajectory stability after inserting a branch at the end node; (a) Before inserting the branch, the trajectory of state evolution and the flow of each link; (b) After inserting the branch, the trajectory of state evolution and the flow of each link
图4. 末端分支链路对ITC模型的路径稳定性影响分析;(a) 加入末端分支链路之前,状态演化路径及各路径通量情况;(b) 加入末端分支链路之后,状态演化路径及各路径通量情况
算出指定节点数目和指定分支位置下的平均W值。总的计算结果如图5所示。
横坐标表示接入分支链路的位置,−1表示未接入分支链路,0表示独立链路,1表示起始节点,以此类推。对于n个节点的网络,横坐标为n − 1就是末端分支链路。从图5中可以看出,随着接入位置的后移,网络W值先降低然后增加。这反映了接入分支链路后的两种作用,接入位置靠前,减小路径重合度,从而降低W值;而接入位置靠后,演化路径会被延长,从而导致W值会显著增加,这一点与理论分析的结果完全吻合。
4. 讨 论
本文以ITC网络模块为基础,研究了调控网络中分支链路的接入位置对路径稳定性影响情况,通过理论推导表明,一方面,在ITC模型的起点或者ITC模型之外接入分支链路,状态演化路径的汇聚时间点被推迟,从而导致路径重叠度降低,进而是W值减少;另一方面,在ITC模型的末端接入分支链路,
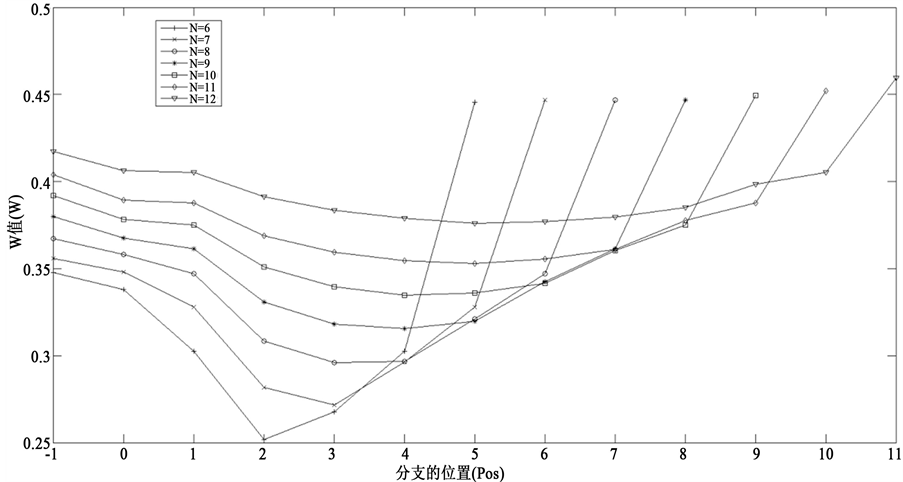
Figure 5. Simulation results of the effect of branch’s position on the trajectory stability. Pos = –1 means there is no branch. Pos = 0 means the independent branch. Pos = 1 - n – 1 indicates that inserting branch from the start node to the end node in succession
图5. 分支链路对路径稳定性的影响模拟分析图。位置为–1表示没有接入分支,位置为0表示接入独立分支,位置为1~ n – 1表示从网络起点开始依次接入分支
状态演化路径会被延长,从而提高了网络的W值。此外,实验模拟的结果也进一步表明在ITC模型的中间节点接入分支链路,其对网络路径稳定性的影响是这两种作用的综合。而且,越靠近起点,W值减少的效果明显,越靠近末端,W值增加的效果明显。本研究结论不仅为解析路径稳定性的结构成因奠定基础,还能够指导合成生物学研究中设计出具有较高路径稳定性的网络结构。